In Biodiverse this can be done using the Run Exclusions dialogue, accessed via the Basedata menu.
The main principle of this dialogue is to allow the user to select some set of properties, of either labels or groups, and then remove them from the basedata. A simple example might be to remove all groups containing five or fewer labels, or labels with ranges exceeding some threshold. More complex queries can also be specified using text matching (labels) or spatial conditions (for groups).
One can also delete labels using the selection menu in the View Labels tab (see this previous post), but this does not apply to groups (unless you transpose the basedata so you can treat the groups as labels, and then transpose them back).
An important point to note is that the Run Exclusions dialogue does not trigger updates in any open View Labels tabs. If you delete labels or groups from a basedata then you will need to close and re-open any View Labels tabs for that basedata to see the changes.
It is also worth noting that the aggregation of labels to groups is a one way process. The original input records cannot be recovered from a basedata object unless a cell size of zero is used (for numeric axis data) or there is only one record per text axis (when text axes are used). It is impractical to store the 10 GB data from the example at the top.
The rest of this blog is just a set of examples. The data are the example data provided with Biodiverse. There are also details in the help system at https://github.com/shawnlaffan/biodiverse/wiki/SampleSession#excluding-data
 |
| The Exclusions dialogue is accessed through the Basedata menu. Exclusions will apply to the currently selected Basedata object. |
 |
| View Labels tab showing the original distribution of data. |
 |
| This example uses a definition query to delete all groups whose centroid falls inside polygons where the state field has a value of 'Tas'. |
 |
| The system gives feedback about how much was deleted. |
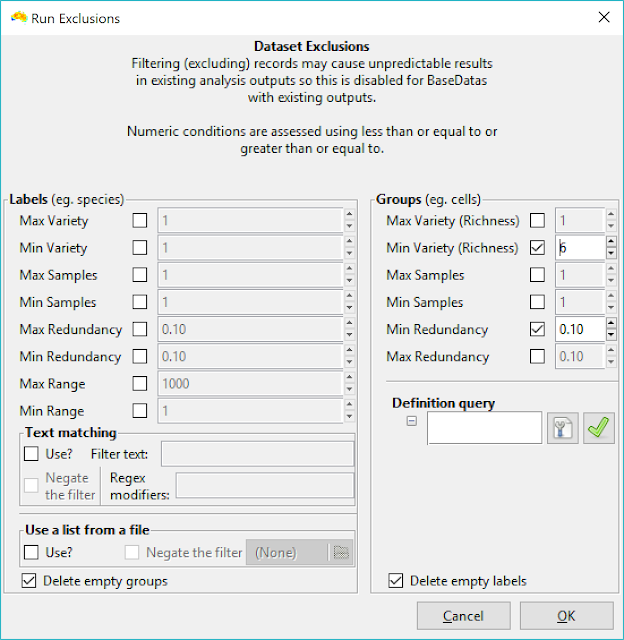 |
| This example will remove all groups with fewer than 6 species, or where the sample redundancy score is less than 0.1. |
 |
| The feedback (based on a fresh copy of the basedata) |
 |
| Not much is left in this case... |
 |
| And the feedback. Only two labels are deleted, as the only matching labels in this basedata are Genus:sp12 and Genus:sp22. |
That's about it, really. If you have questions then they can be posted to the Biodiverse user group, or to the google plus page (until Google shuts that service down).
Shawn Laffan
28-Feb-2019
--------
For more details about Biodiverse, see http://shawnlaffan.github.io/biodiverse
To see what Biodiverse has been used for, see https://github.com/shawnlaffan/biodiverse/wiki/PublicationsList
You can also join the Biodiverse-users mailing list at http://groups.google.com/group/Biodiverse-users or follow the google plus page: https://plus.google.com/+BiodiverseSoftware


No comments:
Post a Comment
Note: only a member of this blog may post a comment.