This makes it easier to use the regions in geospatial analyses, for example as conditions in spatial analyses in Biodiverse. Perhaps the best example is that you can generate a regionalisation using one data set, and then assess its values for one or more other data sets. See for example González-Orozco et al. (2013, 2014a and 2014b).
The option is available via the export menu for cluster and region grower analyses. The options are all the usual ones, although that is not many in this case.
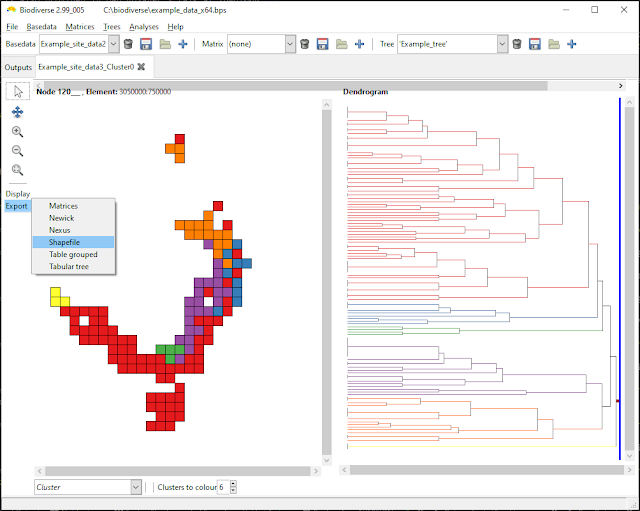 |
| The option is chosen from the export menu for a cluster (or region grower) analysis. |
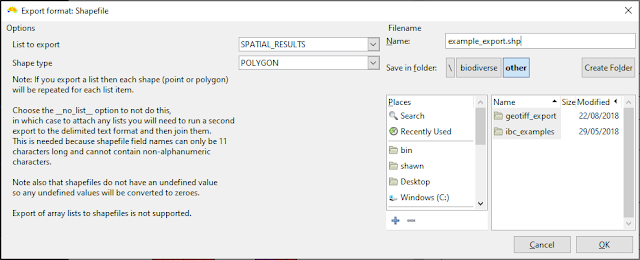 |
| There are not that many choices, but Shape type can be exported as POLYGON or POINT geometries (shapes). You can attach lists of results, or choose to only export the geometries themselves. |
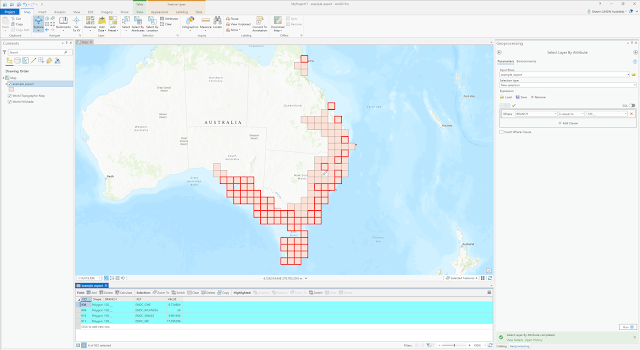 |
| And here the component polygons that form the polygons for branch 120___ are highlighted in red. |
There are three points to watch for.
1. If you export a list with the data, then each polygon or point feature is repeated for each item in the list. This is due to shapefile field names being limited to 11 characters, which is far too short for many of the list entries in Biodiverse. (Support for the GeoPackage format is in the works to obviate this issue).
2. The exported features (geometries) are multipolygon or multipoint. This means that each geometry comprises one or more geometries, with each internal branch including all polygons of its terminal branches. The internal boundaries between polygons are not dissolved, so you will see all the component polygons, but any GIS will support a dissolve operation. If you are wondering what multipolygons and multipoints are, then Wikipedia has a decent explanation as part of the Well Known Text entry.
3. Remember that Biodiverse knows nothing about coordinate systems and map projections (agnostic might be a better way of putting it). You will need to define the coordinate system of the output file yourself using a GIS or other geospatial tool.
Shawn Laffan
12-August-2019
If you want to try this out before version 3 is released then the 2.99_005 development release can be accessed through the downloads page at https://github.com/shawnlaffan/biodiverse/wiki/Downloads
--------
For more details about Biodiverse, see http://shawnlaffan.github.io/biodiverse
To see what Biodiverse has been used for, see https://github.com/shawnlaffan/biodiverse/wiki/PublicationsList
You can also join the Biodiverse-users mailing list at http://groups.google.com/group/Biodiverse-users
No comments:
Post a Comment
Note: only a member of this blog may post a comment.