Biodiverse supports a range of randomisations to assess significance of analysis results. Most use cases in the published literature use the rand_structured algorithm, which is explained in this post, but several common algorithms are supported.
One of the design principles of Biodiverse is to give the user choice. To that end, the curveball algorithm is available from version 5.
The publication describing Curveball is Strona et al. (2014). The name is derived from a baseball card trading card pastime popular in North America.
The curveball algorithm is applied to a data set of items (species, genera, words, or some other set of identifiers). In the common biodiversity case this is a sites by species matrix, transformed to a list of lists, e.g. a list of site lists, where each site list comprises its species (or vice versa). These lists can be considered as sets. At each iteration, two lists (sets of items) are randomly selected. Any items found in both sets are ignored. The rest can be swapped between the two sets, with the number swapped limited by the smaller number of unique items in the two sets to ensure after swapping that each set retains the same number of items it started with. As an example, consider the case where set 1 has ten items, set 2 has eight, and there are six common items found in both lists. This means two items can be swapped between the two lists.
The general formula for the number of possible swaps at an iteration is (min (|A|,|B|) - |A ∩ B|), where A and B are the two sets being considered, and the pipes || denote the lengths of the sets (the numbers of items they contain). If one prefers to think in terms of dissimilarity measures where a is the number of shared items, b the number unique to set 1 and c the number unique to set 2, then the formula is (min (b,c)). Purely as an aside, this is also part of the denominator in Simpson's dissimilarity index.
The curveball algorithm is related to the independent swaps algorithm. The chief advantage of curveball over independent swaps is that, because it swaps as many items as it can at each iteration, it converges on a randomised result much faster. Curveball also avoids the main pitfall of the independent swaps algorithm where a pair can be selected that cannot be swapped, thus "wasting" an iteration (swap attempt).
Curveball does, however, have the same issue that independent swaps has in that the user needs to specify the number of iterations over which swaps will be attempted. Too few and the resulting matrix will not be sufficiently random. Too many and time will be "wasted". This is addressed in Biodiverse by optionally tracking which of the original matrix entries have been swapped, and stopping when all have been done (the stop_on_all_swapped parameter). This has some overhead in the tracking but generally this should be balanced by the time saved by running fewer iterations overall. For those interested, the default number of swaps is the same as for the independent swaps algorithm, which is twice the number of non-zero matrix entries (twice the sum of the lengths of all lists).
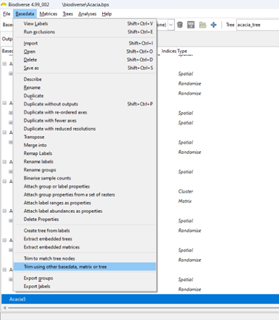
Accessing the curveball algorithm in Biodiverse is the same as for any of the randomisations. Open the Randomisation tab, select rand_curveball as the randomise function, select the number of randomisation iterations and any other algorithm specific parameters, then press Go (see image below). The results are in the same format as always (e.g. see here, here and here).
Since it is just another algorithm, all the common options are available (another new change in version 5 is that more options are available across all algorithms in the GUI - see issue 946). Users can define regions that are randomised separately before reassembly for analysis, including some that are not to be randomised. One can also add some of the randomised results to the project to inspect them.
In terms of speed, curveball is faster than rand_structured. This is largely due to there being less book-keeping required. However, as with independent swaps, curveball can only be applied on a per-cell basis. It does not extend to spatially structured randomisations like rand_structured does (one could ensure swap candidates come from within some local neighbourhood, but this is a different model to something like a diffusion process or a random walk. Update 20241109: This has been implemented and will be available in V5).
 |
| All that is needed to run the curveball algorithm is to choose rand_curveball as the "Randomise function". Other parameters are set as usual. |
And that's pretty much it for the description. If you want to read more randomisation related blog posts then check out the posts tagged with the randomisation label.
Shawn Laffan
05-Nov-2024
For more details about Biodiverse, see http://shawnlaffan.github.io/biodiverse/
For a list of some of the analyses Biodiverse has been used for, see https://github.com/shawnlaffan/biodiverse/wiki/PublicationsList
You can also join the Biodiverse-users mailing list at https://groups.google.com/group/Biodiverse-users or start a discussion at https://github.com/shawnlaffan/biodiverse/discussions